1. Study the effects of long-range and high forces generated by cancer cells on tumor microenvironment.
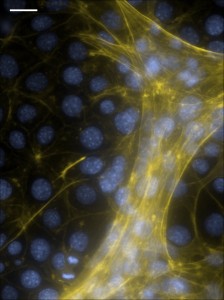
Some types of cancer cells can generate higher forces compared to their normal counterparts. The high forces can be transmitted over distance at the order of hundreds of microns. There can be two possible effects: (a) activation of mechanosignaling pathways in cells that are not in the immediate proximity of the cancer cells; and (b) mechanical remodeling of the microenvironment as a result of reorganization of fibrillar ECM proteins (see movie below) and subsequent changes in the diffusion rate of secreted substances such as cytokines, microRNAs and exosomes.
We are looking into how these effects contribute to cancer pathology.
2. Construct physiologically representative organoid systems by mimicking in vivo biophysical characteristics.
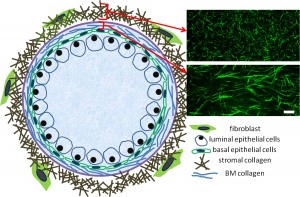
In order to have a better representation of physiological conditions, we are interested in developing an organoid system that takes into account the in vivo biophysical factors such as ECM organization, rigidity, viscosity of the interstitial fluid, and so on. Hopefully the pharmaceutical test results derived from such organoid system can better predict the outcomes of clinical trials than the currently prevailing cell culture model systems do.
3. Innovative instrumentation for tissue/cell mechanics studies.
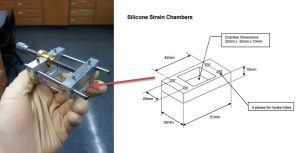
We design and build instruments allowing us to studying the responses of cells and/or tissues to mechanical stimuli or lack thereof. One example is the universally fitted, real-time imaging capable stretcher. We can directly monitor fast cell responses at high resolution using the stretcher.
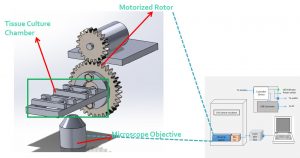
We also build microgravity simulators where with a one- or two-axis rotation, cultured tissues are accelerated until reaching terminal velocity at which the gravitational force is counterbalanced by the hydrodynamic forces. The simulator has the capacity to measure cardiac tissue contractility and apply pulsative tensile forces to simulate heart beats. It will help us understand how the heart is impacted by the microgravity in the space and whether we can rehabilitate the cardiac functions by externally applying forces longitudinally.
We welcome inquiries from potential collaborators for possible applications of our instruments. Please email Yun Chen ([email protected]) for details.
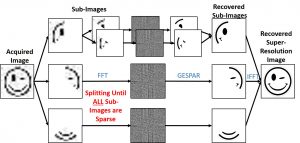
4. New imaging platforms and computational super-resolution.
Recently developed tools in biochemistry, molecular biology, biophysics contribute greatly to advancing our understanding how molecules interact with each other in healthy and abnormal cells. However, sometimes what we know at the molecular level cannot be linearly scaled up to the tissue level; and such “not seeing the wood for the trees” issue arises from the fact that heterogeneity of the tissue is not accounted for. In order to understand the inter-scale relationship between molecules, cells and tissues in homeostasis and pathology, we will address the issue by developing a new imaging platform and corresponding probes to survey the tissue heterogeneity at single-molecule level by combining coherent diffraction and compressed sensing for image reconstruction achieving sub-wavelength resolution.
5. Map the heterogeneity in tumor microenvironment.
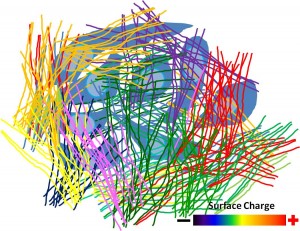
Tumor heterogeneity presents major problems limiting the efficacy of targeted therapies. There is a vast body of evidence that breast cancer is a heterogeneous disease . It is believed that tumor microenvironment plays an important role in driving tumor heterogeneity, though no detailed mechanism has been proposed and proven. By visually examining the tissues from breast cancer patients, it was noted that stroma surrounding cancer cells bearing different mutations also differs in texture and abundance of collagen fibers. We are systematically testing if the heterogeneity in microenvironment, in terms of ECM composition, rigidity, diffusion rate of macromolecules, surface charge density, etc., contributes to tumor heterogeneity.
See more details at https://scholar.google.com/citations?hl=en&user=0jZf7TMAAAAJ&view_op=list_works&sortby=pubdate